L4 phytoplankton time-series
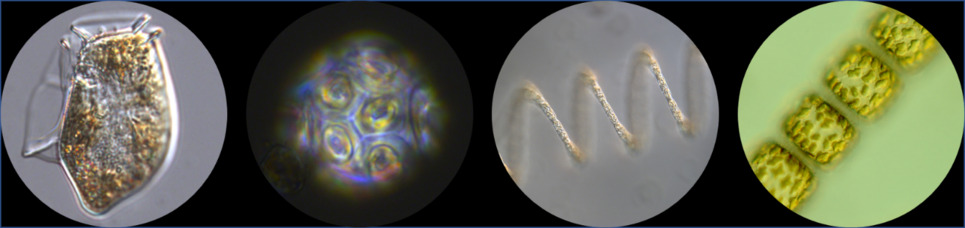
The Station L4 (50°15.0'N; 4°13.0'W) phytoplankton time-series (Widdicombe et al 2010) contains taxa-specific and total functional group abundance (cells mL-1) and biomass (mgC m-3) data.
Samples have been collected weekly (weather permitting) from the 10m depth since October 1992, whereby paired 200 mL water samples are fixed in acid Lugol's iodine (Throndsen, 1978) for enumerating all phytoplankton cells > 2µm and neutral formaldehyde for enumerating coccolithophores. Samples are analysed at Plymouth Marine Laboratory according to the Utermohl counting technique (Utermohl, 1958) and since 2016, these methods follow the British and European standard document "Water quality - Guidance standard for routine microscopic surveys of phytoplankton using inverted microscopy (Utermohl technique)" (BS EN 15204:2006).
Individual taxa are grouped into Diatoms, Dinoflagellates, Coccolithophores, Flagellates, Phaeocytis and Ciliates and taxa-specific cell biovolumes (e.g. Olenina et al 2006) are converted to carbon (pgC cell-1) using the formulae of Menden-Deuer and Lessard (2000). Two analysts have generated the data since 1992 and in 2005 more ciliates were identified to species level hence the increase in the number of taxa. Further information on individual biovolume and carbon values, identification consistency and data quality issues are found in the taxa-specific metadata file.
For queries and access to the dataset please contact Claire Widdicombe (clst@pml.ac.uk).
We view it as essential for all users of L4 plankton data to establish and maintain contact with the nominated current data originators as well as fully consulting the metadata. While not impinging on free data access, this ensures that this large, species-rich but slightly complex species database is being used in the correct way, and any potential issues with the data are clarified. Furthermore, a proper dialogue with these local experts on the time series will enable where appropriate the most recent sampling timepoints to be used.
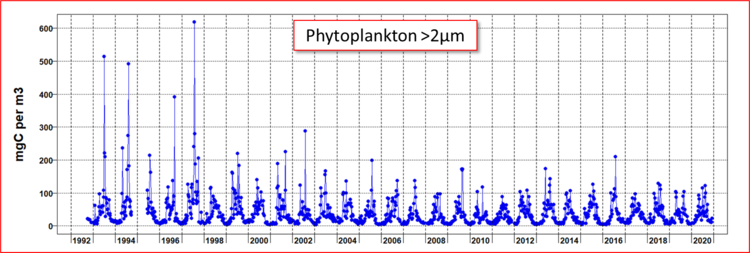
Phytoplankton carbon at L4
References
British Standard (EN15204:2006) Water quality - Guidance standard on the enumeration of phytoplankton using inverted microscopy (Utermohl technique) 42 pp.
Olenina, I., Hajdu, S., Edler, L., Andersson, A., Wasmund, N., Busch, S., Gobel, J., et al. 2006. Biovolumes and size-classes of phytoplankton in the Baltic Sea. HELCOM Balt. Sea Environ. Proc., 166: 144 pp.
Throndsen, J. 1978. Preservation and storage. In Sournia, A. (ed.), Phytoplankton manual. UNESCO, Paris. pp 69-74.
Menden-Deuer, S., Lessard, E.J., (2000), Carbon to volume relationships for dinoflagellates, diatoms, and other protist plankton, Limnology and Oceanography, 3, doi:10.4319/lo.2000.45.3.0569.
Throndsen, J. (1978). Preservation and storage. In: Sournia, A. (ed ) Phytoplankton manual. UNESCO, Paris, p. 69-74
Utermohl, H. 1958. Vervollkommung der quantitativen Phytoplankton-Methodik. Mitt. Int. Ver. Limnol., 9:1-38 Widdicombe, C. E., Eloire, D., Harbour, D., Harris, R. P. and Somerfield, P. J. 2010. Long-term phytoplankton community dynamics in the Western English Channel. J. Plankton Res., 32: 643-655
Widdicombe, C. E., Eloire, D., Harbour, D., Harris, R. P. and Somerfield, P. J. (2010) Long-term phytoplankton community dynamics in the Western English Channel. J. Plankton Res., 32: 643-655
Widdicombe C.E.; Harbour D.(2021). Phytoplankton taxonomic abundance and biomass time-series at Plymouth Station L4 in the Western English Channel, 1992-2020. NERC EDS British Oceanographic Data Centre NOC. doi:10/grks.
 Western Channel Observatory
Western Channel Observatory